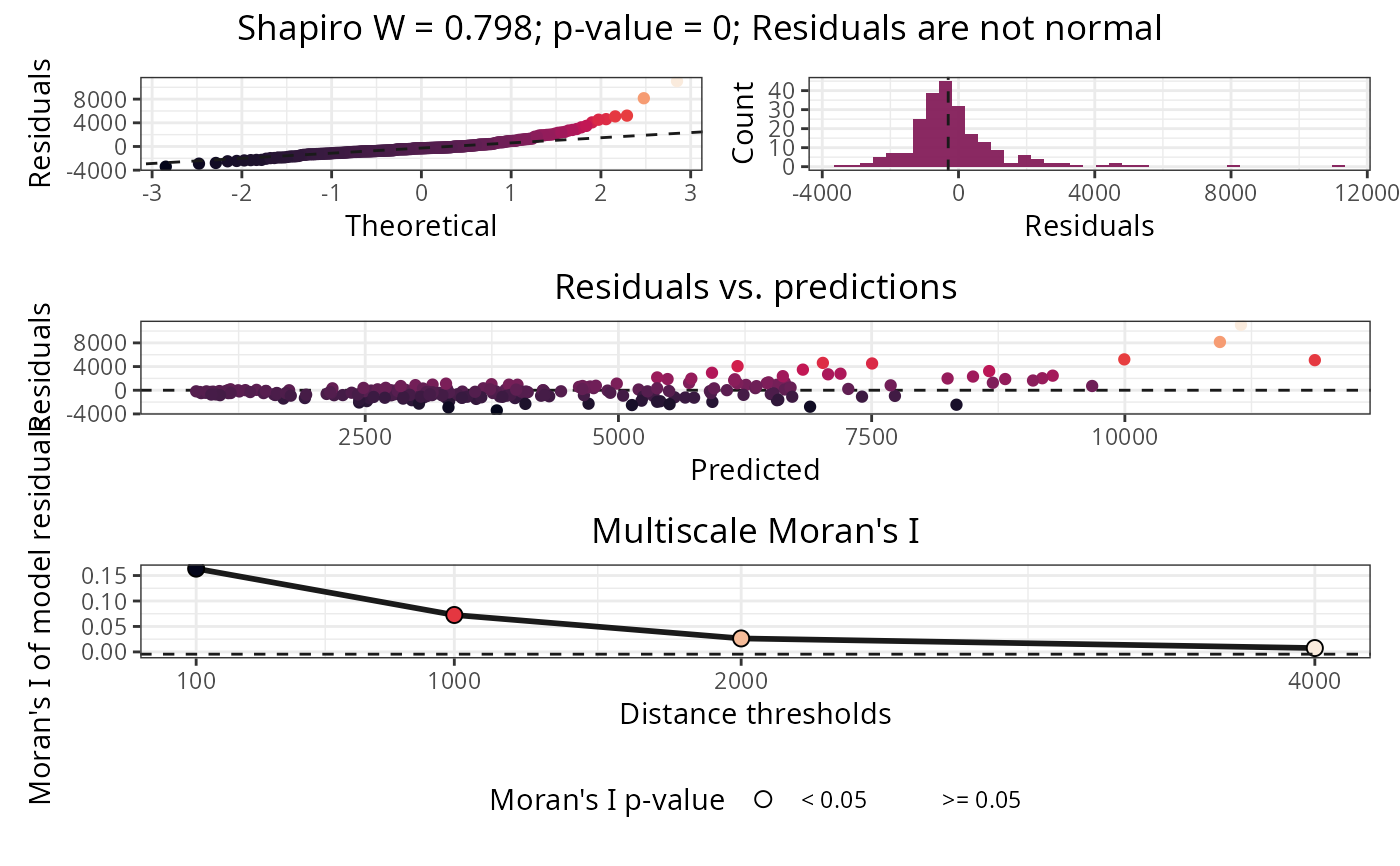
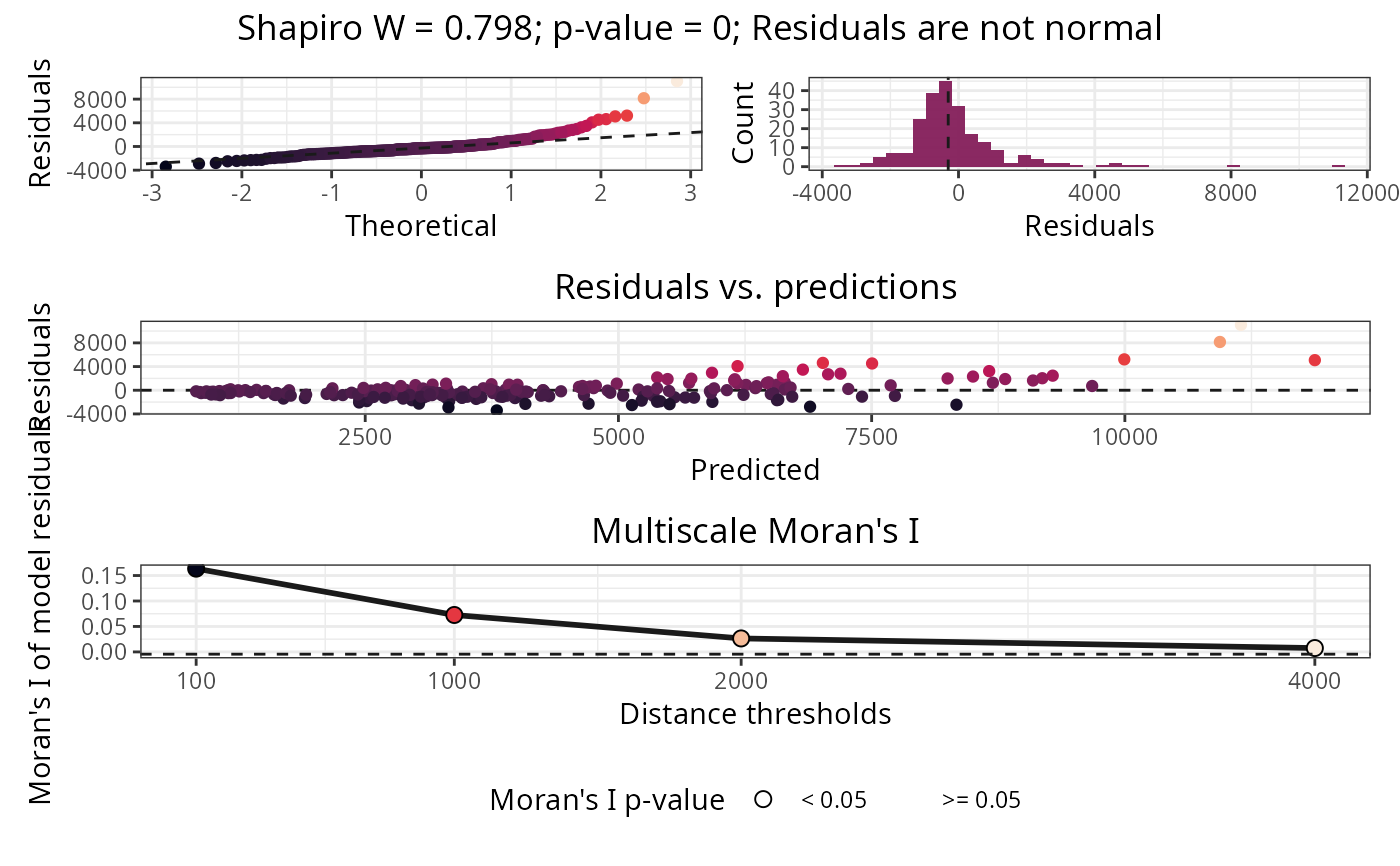
Plots normality and autocorrelation tests of model residuals.
Arguments
- model
A model produced by
rf(),rf_repeat(), orrf_spatial().- point.color
Colors of the plotted points. Can be a single color name (e.g. "red4"), a character vector with hexadecimal codes (e.g. "#440154FF" "#21908CFF" "#FDE725FF"), or function generating a palette (e.g.
viridis::viridis(100)). Default:viridis::viridis(100, option = "F")- line.color
Character string, color of the line produced by
ggplot2::geom_smooth(). Default:"gray30"- fill.color
Character string, fill color of the bars produced by
ggplot2::geom_histogram(). Default:viridis::viridis(4, option = "F", alpha = 0.95 )[2]- option
(argument of
plot_moran()) Integer, type of plot. If1(default) a line plot with Moran's I and p-values across distance thresholds is returned. If2, scatterplots of residuals versus lagged residuals per distance threshold and their corresponding slopes are returned. In models fitted withrf_repeat(), the residuals and lags of the residuals are computed from the median residuals across repetitions. Option2is disabled ifxis a data frame generated bymoran().- ncol
(argument of
plot_moran()) Number of columns of the Moran's I plot ifoption = 2.- verbose
Logical, if
TRUE, the resulting plot is printed, Default:TRUE
Examples
data(plants_rf)
plot_residuals_diagnostics(plants_rf)